Name: Yogindra Raghav
Type: User
Company: University of Virginia (UVA) Medicine
Bio: Aspiring Scientist, Author, Philosopher. Computational Biology PhD Student @ UVA Medicine
Location: Charlottesville, VA
Blog: https://www.linkedin.com/in/YogiOnBioinformatics/
Yogindra Raghav's Projects
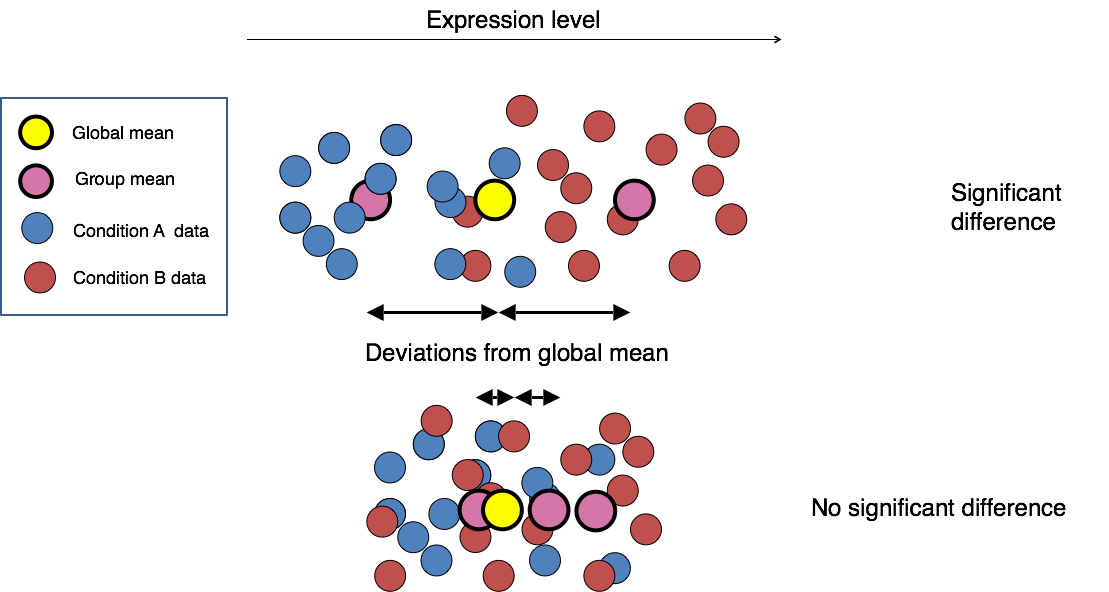
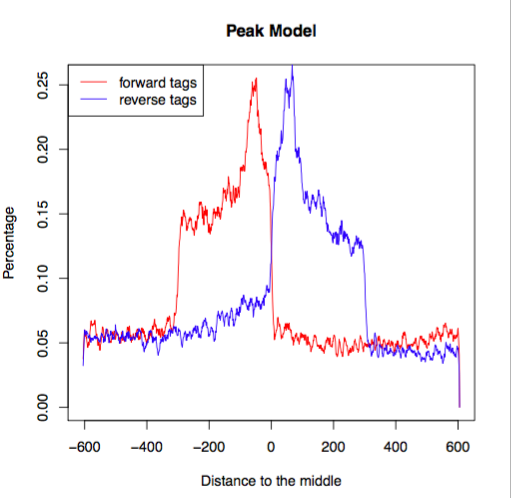
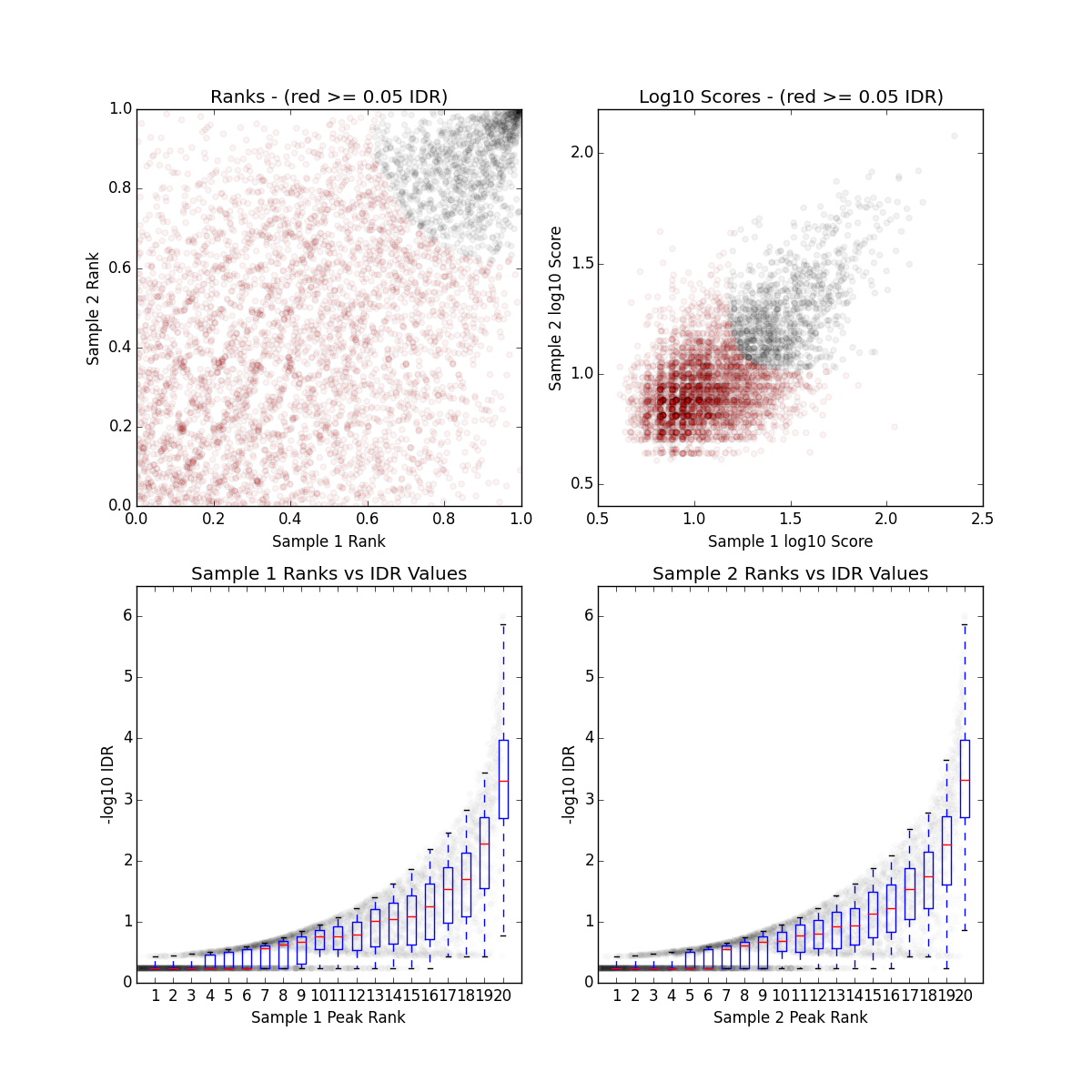
Determining reproducibility of Epigenomic signal for Answer ALS
Projects and Essays from Algorithm Implementation (CS 1501) at University of Pittsburgh
Handwritten work & R code related to University of Virginia's MATH 3350 ( Applied Linear Algebra)
Biplots, Volcano plots, PCA plots, Heatmaps and more Computational Genomics data created and visualized during University of Pittsburgh course, Computational Biology (BIOSC1540), with Dr. Miler Lee
Work done during rotation with Coleen McNamara & Stefan Bekiranov in UVA BIMS PhD program
Worked with Dr. Shandong Wu at University of Pittsburgh to use software to improve outcomes for breast cancer risk detection.
Published work of mine in Pitt Biological Sciences Advising Blog about using Bioinformatics to predict Ligand-Protein interactions.
Challenge to efficiently BLAST files (FASTA file & quality scores) and report key metrics about FASTA back to user.
Project in the Durrant Lab at UPitt that wanted to re-use code from a previous neural network ligand-protein interaction software to extract features for ML
Independent project I undertook to perform a full ChIP-Seq analysis of the transcription factor Nanog in Zebrafish embryos.
Containers that I use in my work.
Links to other repositories containing all the work I did as a Computational Biology Research Assistant in the Dr. Jacob D. Durrant Lab of Computational Drug Discovery, University of Pittsburgh.
Description of work done at Merck pharmaceutical company in the summer of 2018 as a Computational Drug Discovery Intern at West Point, PA. Information excludes all proprietary information belonging to Merck & Co.
Work done during rotation with Ani Manichaikul during UVA PhD program
Destruction of Aqueous Microfibers Enzymatically (DAME): A novel Synthetic Biology systems approach to destroying Polyester-based Microplastic fibers
A repository containing examples of Data Structure Implementations that I did during my University of Pittsburgh course CS445: Data Structures-Abstractions and Implementations
Worked with fellow peers at University of Pittsburgh under supervision of Dr. Junshu Bao, Department of Statistics, University of Pittsburgh, to do a Differential Gene Expression Analysis on a Dexamethasone treatment data set and incorporated Machine Learning into project.
Dotfiles + config files that I use in my personal and professional work.
Independent Data Science project undertaken to investigate drug usage among the Baby Boomer generation in the recent past.
Validating the first human iPSC-derived Cortical Neuron ALS model via systematic Epigenomic analysis.
Integration of ATAC-Seq and multi-assay ChIP-Seq data using IDEAS (Hidden Markov Model) approach.Multi-Epigenomic state annotation with CHDI NeuroLINCS Epigenomics data using the tool "IDEAS"
Joint Genotyping in MIT Fraenkel Lab with ALS Genomics + 1000 Genomes Data
Research project undertaken in Computational Genomics at the University of Pittsburgh. with Dr. Miler Lee and students to understand Maternal-to-Zygotic transition in Xenopus laevis
Examples of Molecular Phylogenomics figures created from self-taught usage of MEGA-X software.
Ordinal Logistic Regression with ElasticNet Regularization using Multi-Assay Epigenomics Data from CHDI NeuroLINCS Consortium.
This repository contains examples of non-scientific writing that I have done. This may be useful for those wanting to know about my writing style and ability since it is useful in the scientific world.
Work done for University of Pittsburgh course "Principles of Data Science" (STAT 1261) with Dr. Junshu Bao in Fall semester of 2018.
Protein Virtual Reality is a University of Pittsburgh project with Dr. Jacob D. Durrant that is the world's first anatomical and protein visualization VR program tailored for collegiate-biology courses.
Common resource files necessary for Computational Biology research.
Contains publications that I wrote for the PittPulse during my time as a Senior Computational Biology Correspondent.