Here are some elapsed times (in s) for 5 implementations (of course, these numbers do not characterized the languages but only particular implementations in some languages).
| # particles | Py | C++ | Fortran | Julia | Rust |
|---|---|---|---|---|---|
| 1024 | 30 | 55 | 41 | 45 | 34 |
| 2048 | 124 | 231 | 166 | 173 | 137 |
| 16384 | 7220 | 14640 | 10914 | 11100 | ? |
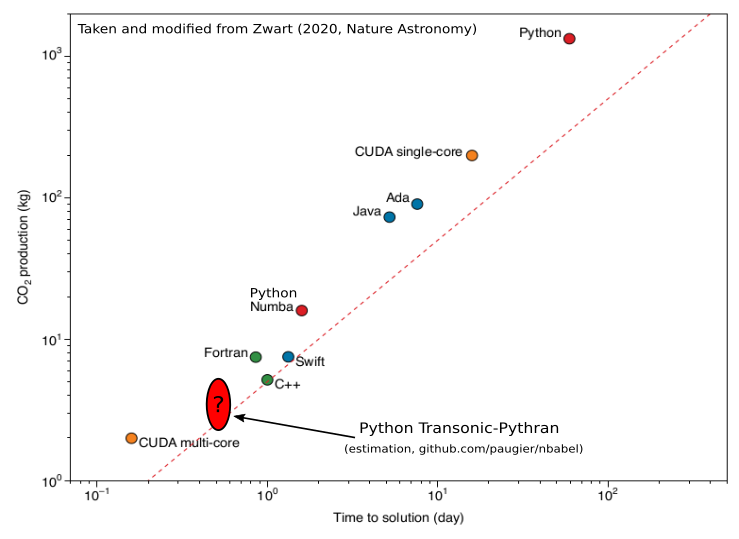
The implementations in C++, Fortran and Julia come from https://www.nbabel.org/ and have recently been used in an article published in Nature Astronomy (Zwart, 2020). The implementation in Python-Numpy is very simple, but uses Transonic and Pythran (>=0.9.8).
To run these benchmarks, go into the different directories and run make bench1k or make bench2k.
To give an idea of what it gives compared to the figure published in Nature Astronomy:
Note: these benchmarks are run sequentially with a Intel(R) Core(TM) i5-8400 CPU @ 2.80GHz.
Note 2: With Numba, the elapsed times are 44 s, 153 s and 11490 s, respectively. This is approximately 20% faster than the C++ implementation.
Note 3: With PyPy, a pure Python implementation (bench_pypy_Point.py) runs for 1024 particles in 151 s, i.e. only 3 times slower than the C++ implementation (compared to ~50 times slower as shown in the figure taken from Zwart, 2020).
Note 4: The directory "julia" contains some more advanced and faster implementations. The sequential optimized Julia implementation runs on my PC in 27.2 s, 104.4 s and 8900 s, respectively (i.e. +- 20% compared to our fast and simple Python implementation).
Note 5: From the high level Numpy implementation
(bench_numpy_highlevel.py), if one (i) adds an import from transonic import jit and (ii) decorates the function loop with @jit, the case for 1024
particles runs in 136 s (2.5 times slower than the C++ implementation).
We can also compare different solutions in Python. Since some solutions are very slow, we need to compare on a much smaller problem (only 128 particles). Here are the elapsed times (in s):
| Transonic-Pythran | Numba | High-level Numpy | PyPy OOP | PyPy lists |
|---|---|---|---|---|
| 0.48 | 0.87 | 686 | 2.6 | 4.3 |
For comparison, we have for this case {"c++": 0.85, "Fortran": 0.62, "Julia": 2.57}.
Note that just adding from transonic import jit to the simple high-level
Numpy code and then decorating the function loop with @jit, the elapsed
time decreases to 2.1 s (a ~ x300 speedup!, with Pythran 0.9.8).