The purpose of this git repo is to share the scripts, files and tests of the tools I developed during the period of April 5 until June 1.
Having the scientists with no prior coding knowledge in mind, these UIs were conceived in the direction of making the analysis steps easier to launch, verify and visualize.
These tools are hosted on the CIPHE server as will be shown in the examples and tutorials available inside the files. They are all based on R and R Shiny.
As a result, the main way to test the different tools is to connect to the CIPHE network.
Alternatively, I can help in providing support in modifying the scripts in order to test them in local for people that are interested.
The tutorials for each of the applications is provided in their corresponding folders, this README is merely an introduction.
The main goal of the internship in the beginning was just the conception of a user-friendly graphical interface for the infinityFlow R package. Evidently, we ended up doing more than that by creating three different tools that can each be used on their own or be related to each other to create a complete pipeline that satisfies the purpose of phenotyping the immune cells.
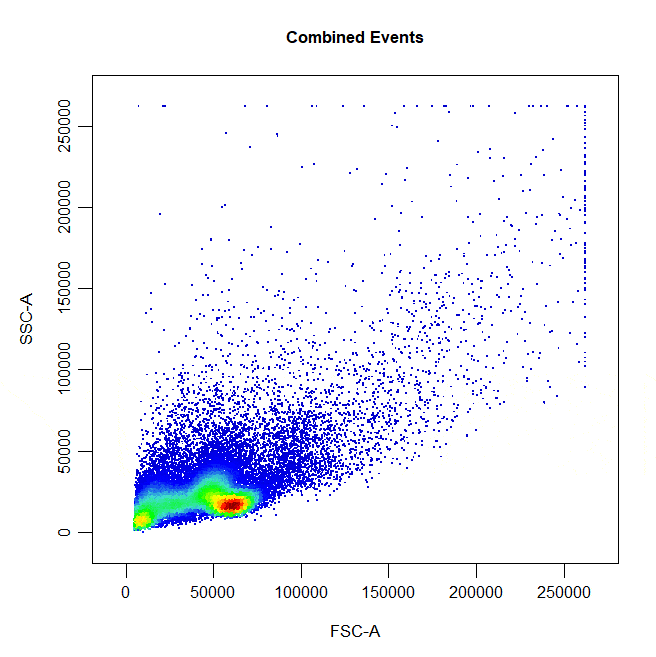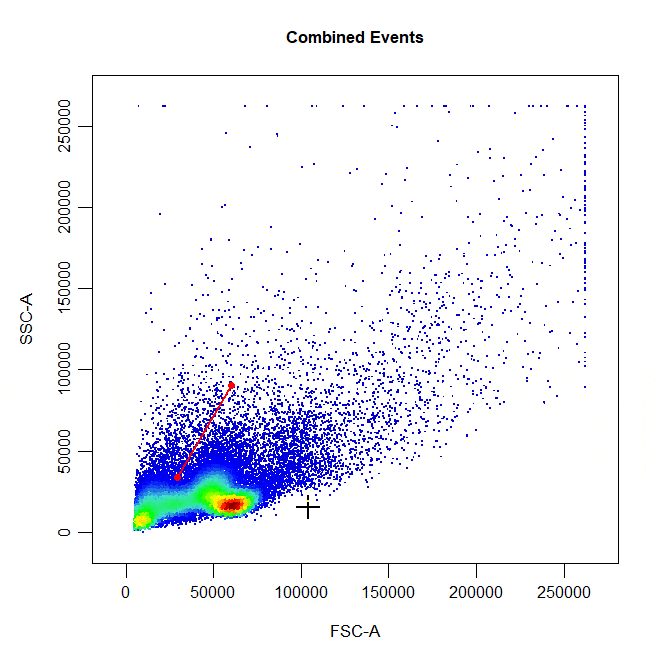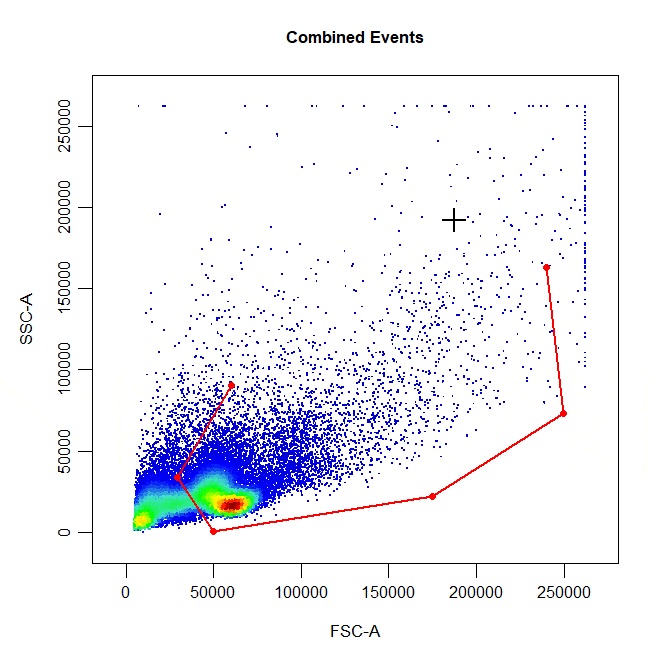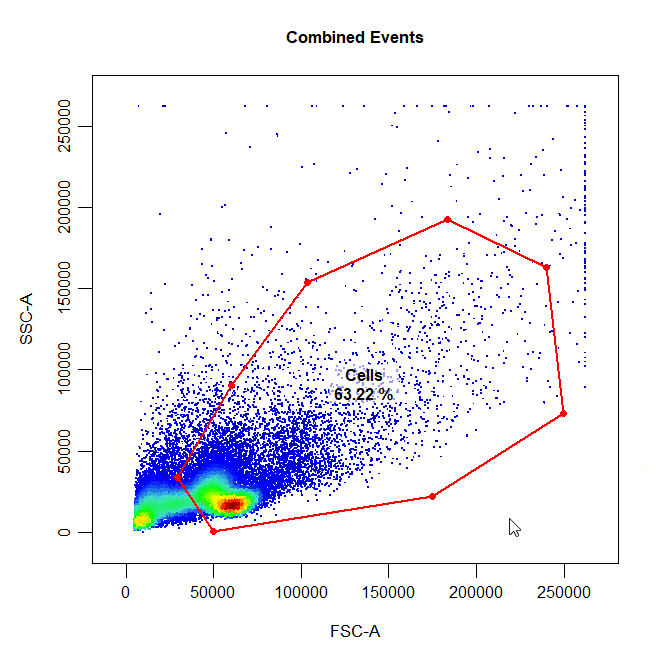
State of the art gating tool for flow cytometry data based on (https://github.com/DillonHammill/CytoExploreR). This tool not only accepts different type of formats of gating software, it also indexes the files and annotates the events.
A mini-tool that fills the purpose of choosing the same number of cells for all the populations for further downstream analysis. Used so the machine learning model in infinity Flow will capture the characteristics of the rare populations.
Also, equally sampling files can be useful for clustering and adapting a supervised approach face-to-face rare populations.
XGBoost-based machine learning approach that reduces the noise-to-signal ratio of flow cytometry data based on (https://github.com/ebecht/infinityFlow). Additional tools were added to generate statistics and visualize the plots.
A collection of R scripts that were used on the real dataset mainly to verify the results. Some of these scripts have a more specific role of visualizing or comparing two files.
The scripts that were used in the first tool of gating that was used in CIPHE, I also contributed to this tool before switchin to a CytoExploreR based tool.
Please refer to https://github.com/GeorgeAlehandro/cipheinfinityFlow and https://github.com/GeorgeAlehandro/cipheCytoExploreR to oberve the modifications that I made over the two main packages (with all rights reserved to the original creators) in order to ensure a compatibility with R Shiny UI.
Might also be interesting to take a look at both of these issues:
A list of all the packages and dependencies used to run the tools are available in https://github.com/GeorgeAlehandro/CIPHE_internship/blob/main/prerequisites.csv
Check CIPHE Gate and CIPHE Infinity for a step-by-step tutorial for each of the tools.
Also, if you would like to take a look at my M1 internship report paper, I can provide it.
I would like to thank Mr. Etienne Becht and Mr. Dillon Hammil for their support.
I also thank a lot another contributor for the tools, even without coding, Dr. Hervé Luche for his guidance and continuous feedback.