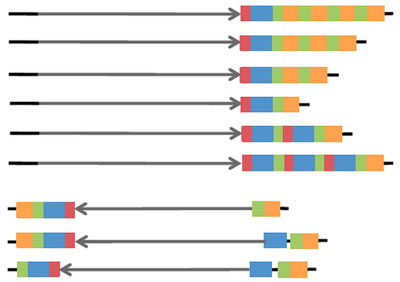
A protocol to impute 17q21.32 haplotypes from surrounding genotypes computed from genotype array or whole genome sequence data using the reference panel from:
Boettger L., McCarroll S., et al. Structural haplotypes and recent
evolution of the human 17q21.31 region. Nat. Genet. 44, 881–885 (2012)
This reference panel for HapMap samples was generated using droplet digital PCR as explained here. For any feedback, send an email to [email protected] or [email protected]
Install basic tools (Debian/Ubuntu specific):
sudo apt install wget gzip samtools bcftools plink1.9 openjdk-11-jre-headless
Preparation steps
mkdir -p $HOME/res
Download Beagle binary
wget -P $HOME/res/ https://faculty.washington.edu/browning/beagle/beagle.25Nov19.28d.jar
Download reference panels
wget -P $HOME/res/ https://personal.broadinstitute.org/giulio/panels/chr17q21_haplotypes_{hm3,1kg}.GRCh3{7,8}.vcf.gz
Run imputation using Beagle
vcf="..."
out="..."
sfx="1kg" # sfx="hm3"
build=38 # build=37
declare -A reg=( ["37"]="17:43165384-45781599" ["38"]="chr17:45088016-47704233" )
bcftools view --no-version "$vcf" -r ${reg[$build]} | \
java -Xmx8g -jar $HOME/res/beagle.25Nov19.28d.jar gt=/dev/stdin \
ref=$HOME/res/chr17q21_haplotypes_$sfx.GRCh$build.vcf.gz out="$out" \
map=<(bcftools query -f "%CHROM\t%POS\n" $HOME/res/chr17q21_haplotypes_$sfx.GRCh$build.vcf.gz | \
awk '{print $1"\t.\t"$2/1e7"\t"$2}')
Extract imputed 17q21.32 alleles into a table
out="..."
build=38 # build=37
declare -A reg=( ["37"]="17:44166101-44166101" ["38"]="chr17:46088735-46088735" )
bcftools index -ft "$out.vcf.gz" && \
bcftools query -f "[%SAMPLE\t%ALT\t%GT\n]" "$out.vcf.gz" -r ${reg[$build]} | tr -d '[<>]' | \
awk -F"\t" -v OFS="\t" '{split($2,a,","); a["0"]="NA"; split($3,b,"|"); \
print $1,a[b[1]],a[b[2]]}' > "$out.tsv"
This section is only in case you want to build the reference panels yourself, you can skip it otherwise
Download GRCh37 human genome reference
wget -O- ftp://ftp.1000genomes.ebi.ac.uk/vol1/ftp/technical/reference/human_g1k_v37.fasta.gz | \
gzip -d > $HOME/res/human_g1k_v37.fasta
samtools faidx $HOME/res/human_g1k_v37.fasta
Download GRCh38 human genome reference
wget -O- ftp://ftp.ncbi.nlm.nih.gov/genomes/all/GCA/000/001/405/GCA_000001405.15_GRCh38/seqs_for_alignment_pipelines.ucsc_ids/GCA_000001405.15_GRCh38_no_alt_analysis_set.fna.gz | \
gzip -d > $HOME/res/GCA_000001405.15_GRCh38_no_alt_analysis_set.fna
samtools faidx $HOME/res/GCA_000001405.15_GRCh38_no_alt_analysis_set.fna
Download 17q21.32 reference panel in Beagle 3 format and an additional file with marker positions for the GRCh37 human genome reference
wget -P $HOME/res/ https://raw.githubusercontent.com/freeseek/impute17q21/master/imputation_cnv_{panel_{hm3,1kg}.bgl,haps_encoded}.txt
Liftover marker positions for the GRCh38 human genome reference
wget http://hgdownload.cse.ucsc.edu/admin/exe/linux.x86_64/liftOver
chmod a+x liftOver
wget http://hgdownload.cse.ucsc.edu/goldenPath/hg19/liftOver/hg19ToHg38.over.chain.gz
for sfx in hm3 1kg; do
tail -n+2 $HOME/res/imputation_cnv_panel_$sfx.bgl.txt | grep -v 17:44166 | \
awk '{print "17\t"substr($2,4)}' > $HOME/res/imputation_cnv_panel_$sfx.GRCh37
tail -n+2 $HOME/res/imputation_cnv_panel_$sfx.bgl.txt | grep -v 17:44166 | \
awk '{print "chr17\t"substr($2,4)-1"\t"substr($2,4)}' | \
./liftOver /dev/stdin hg19ToHg38.over.chain.gz /dev/stdout /dev/stderr | \
awk '{print $1"\t"$3}' > $HOME/res/imputation_cnv_panel_$sfx.GRCh38
done
Generate 17q21.32 reference panels in VCF format for both the GRCh37 and GRCh38 human genome references
declare -A fasta=( ["37"]="$HOME/res/human_g1k_v37.fasta" ["38"]="$HOME/res/GCA_000001405.15_GRCh38_no_alt_analysis_set.fna" )
declare -A chr=( ["37"]="17" ["38"]="chr17" )
declare -A pos=( ["37"]="44166101" ["38"]="46088735" )
for sfx in hm3 1kg; do
grep 17:44166 $HOME/res/imputation_cnv_panel_$sfx.bgl.txt | \
awk 'NR==FNR {if (!($3 in y)) {y[$3]=i; x[i++]=$3} z[$4$5$6$7$8$9$10$11$12$13$14$15]=y[$3]}
NR>FNR {for (j=3; j<=NF; j++) w[j]=w[j]$j}
END {for (i in x) if (i==1) printf "<"x[i]">"; else if (i>0) printf ",<"x[i]">";
printf "\t.\t.\t.\tGT";
for (i in w) printf "\t"z[w[i]]}' $HOME/res/imputation_cnv_haps_encoded.txt - | \
sed 's/\t\([1-9]\)\t\([1-9]\)/\t\1|\2/g' > tmp
for build in 37 38; do
tail -n+2 $HOME/res/imputation_cnv_panel_$sfx.bgl.txt | grep -v 17:44166 | tr ' ' '\t' | cut -f3- | \
paste $HOME/res/imputation_cnv_panel_$sfx.GRCh$build - | \
sed 's/?/-/g;s/\t\([ACGT-]\)\t\([ACGT-]\)/\t\1\2/g' | \
bcftools convert --no-version --tsv2vcf /dev/stdin -c CHROM,POS,AA -f ${fasta[$build]} --samples \
$(head -n1 $HOME/res/imputation_cnv_panel_$sfx.bgl.txt | tr ' ' '\n' | \
tail -n+3 | uniq | tr '\n' ',' | sed 's/,$//') | tr '/' '|' | \
awk -v chr=${chr[$build]} -v pos=${pos[$build]} 'NR==FNR {line=$0}
NR>FNR {if (line && $0!~"^#" && $2>pos) {print chr"\t"pos"\t.\tT\t"line; line=""} print}' tmp - | \
bcftools view --no-version -Oz \
-o $HOME/res/chr17q21_haplotypes_$sfx.GRCh$build.vcf.gz
done
/bin/rm tmp
done
Convert the 17q21.32 and 1000 Genomes project reference panels to plink and then merge to compute consistency
url="ftp://ftp.1000genomes.ebi.ac.uk/vol1/ftp/phase1/analysis_results/integrated_call_sets/ALL.chr17.integrated_phase1_v3.20101123.snps_indels_svs.genotypes.vcf.gz"
bcftools view --no-version $url -r 17:43165384-45781599 | \
awk 'NF==2 {print "##contig=<ID=17,length=81195210>"} {print}' | \
bcftools view --no-version -v snps | \
bcftools annotate --no-version -x ID -I +'%CHROM:%POS:%REF:%ALT' | \
$HOME/bin/plink --vcf /dev/stdin --keep-allele-order --const-fid --make-bed \
--out ALL.chr17.integrated_phase1_v3.20101123.snps_indels_svs.genotypes
for sfx in hm3 1kg; do
bcftools annotate --no-version -x ID -I +'%CHROM:%POS:%REF:%ALT' \
$HOME/res/chr17q21_haplotypes_$sfx.GRCh37.vcf.gz | \
$HOME/bin/plink --vcf /dev/stdin --biallelic-only --keep-allele-order --const-fid --make-bed \
--out chr17q21_haplotypes_$sfx.GRCh37
plink --bfile chr17q21_haplotypes_$sfx.GRCh37 \
--bmerge ALL.chr17.integrated_phase1_v3.20101123.snps_indels_svs.genotypes --merge-mode 6
done
You should get the following results
217071 overlapping calls, 216606 nonmissing in both filesets.
215812 concordant, for a concordance rate of 0.996334.
...
2747672 overlapping calls, 2747672 nonmissing in both filesets.
2741694 concordant, for a concordance rate of 0.997824.