The root-mean-square deviation (RMSD) is calculated, using Kabsch algorithm
(1976) or Quaternion algorithm (1991) for rotation, between two Cartesian
coordinates in either .xyz or .pdb format, resulting in the minimal RMSD.
For more information please read RMSD and Kabsch algorithm.
You have molecule A and B and want to calculate the structural difference between those two. If you just calculate the RMSD straight-forward you might get a too big of a value as seen below. You would need to first recenter the two molecules and then rotate them unto each other to get the true minimal RMSD. This is what this script does.
| No Changes | Re-centered | Rotated |
|---|---|---|
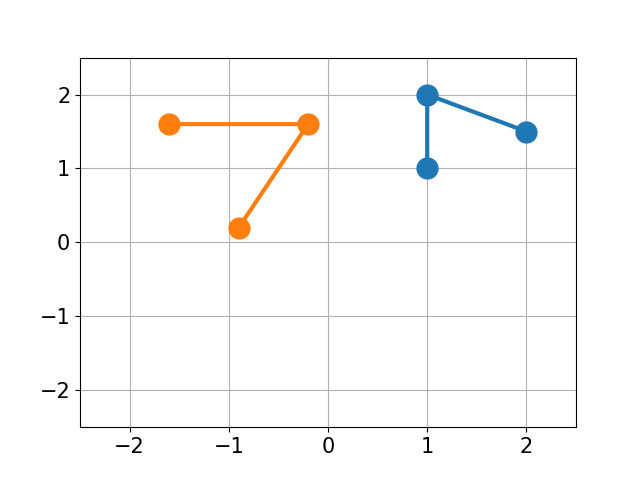 |
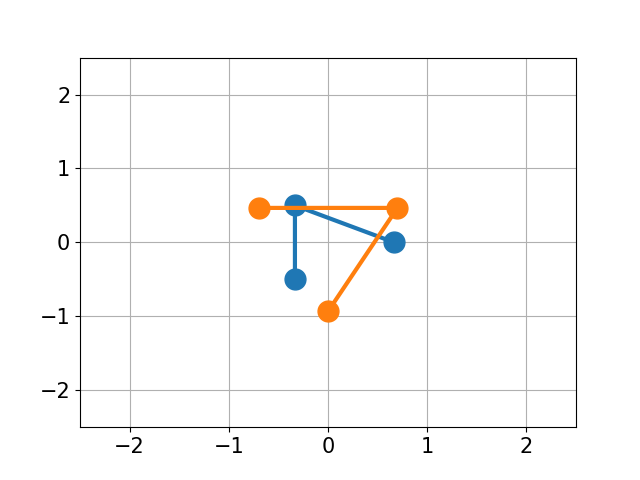 |
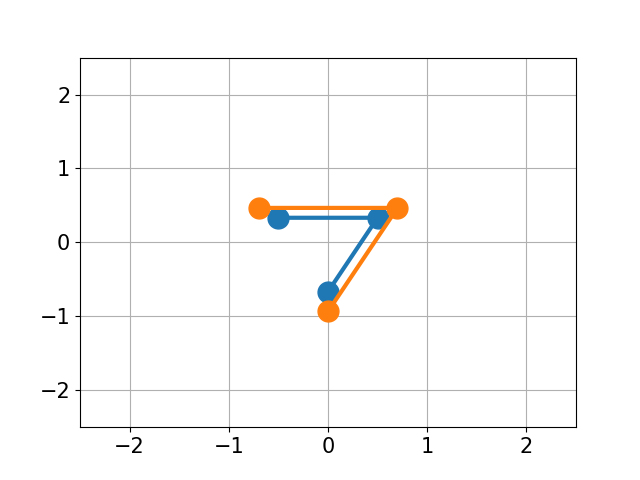 |
| RMSD 2.50 | RMSD 1.07 | RMSD 0.25 |
You can get the package via pip under the name rmsd,
pip install rmsdor download the project from GitHub via
git clone https://github.com/charnley/rmsdThere is only one Python file, so you can also download that and put it in your bin folder.
wget -O calculate_rmsd https://raw.githubusercontent.com/charnley/rmsd/master/rmsd/calculate_rmsd.py
chmod +x calculate_rmsdType calculate_rmsd --help for all the arguments.
Usage is pretty straight forward, call calculate_rmsd with two structures
in either .xyz or .pdb. In this example Ethane has the exact same structure,
but is translated in space, so the RMSD should be zero.
calculate_rmsd examples/ethane.xyz examples/ethane_translate.xyzIt is also possible to ignore all Hydrogens (useful for larger molecules where Hydrogens move around indistinguishable) and output the rotated structure for visual comparison. The output will be in XYZ format.
calculate_rmsd --no-hydrogen --output examples/ethane.xyz examples/ethane_mini.xyzIt is also possible to use RMSD as a library in other scripts:
import rmsd
import numpy as np
P = np.array([[-0.9835 , 1.8109 , -0.0314 ],
[ 0.1268 , 1.8041 , -0.03242],
[-1.4899 , 3.2274 , 0.18102],
[-1.3504 , 1.1535 , 0.78475]])
Q = np.array([[-2.1217 , 4.0933 , 0.12713],
[-1.0113 , 4.0865 , 0.12611],
[-2.628 , 5.5097 , 0.33955],
[-2.4885 , 3.4358 , 0.94328]])
print "RMSD before translation: ", rmsd.kabsch_rmsd(P, Q)
P -= rmsd.centroid(P)
Q -= rmsd.centroid(Q)
print "RMSD after translation: ", rmsd.kabsch_rmsd(P, Q)- Kabsch algorithm:
- Kabsch W., 1976, A solution for the best rotation to relate two sets of vectors, Acta Crystallographica, A32:922-923, doi: http://dx.doi.org/10.1107/S0567739476001873
- Quaternion algorithm:
- Michael W. Walker and Lejun Shao and Richard A. Volz, 1991, Estimating 3-D location parameters using dual number quaternions, CVGIP: Image Understanding, 54:358-367, doi: http://dx.doi.org/10.1016/1049-9660(91)90036-o
- Implementation:
- Calculate RMSD for two XYZ structures, GitHub, http://github.com/charnley/rmsd, <commit hash or version number>
Please cite this project when using it for scientific publications.
Submit issues or pull requests on GitHub.
