README
ETE (Environment for Tree Exploration is a Python programming toolkit that assists in the automated manipulation, analysis and visualization of phylogenetic trees. Clustering trees or any other tree-like data structure are also supported.
ETE is currently developed as a tool for researchers working in phylogenetics and genomics. ETE provides specialized tools to reconstruct, compare and visualize phylogenetic trees. If you use ETE for a published work, please cite:
Jaime Huerta-Cepas, François Serra and Peer Bork. "ETE 3: Reconstruction, analysis and visualization of phylogenomic data." Mol Biol Evol (2016) doi: 10.1093/molbev/msw046
Install/Documentation
- The official web site of ETE is http://etetoolkit.org. Downloading instructions and further documentation can be found there.
- News and announcements are usually posted on twitter: http://twitter.com/etetoolkit
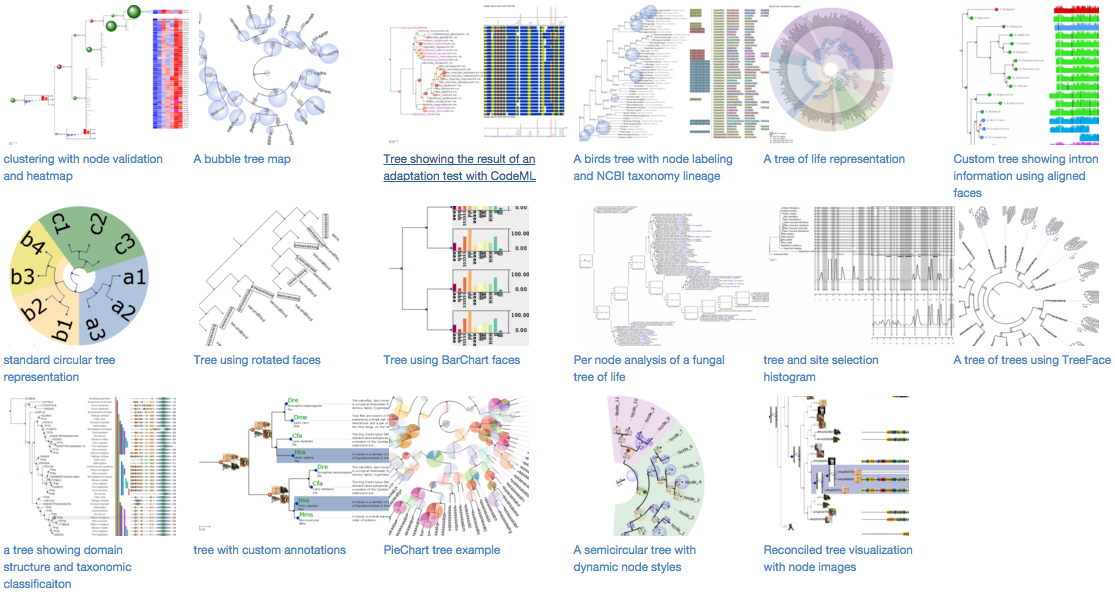
Gallery of examples
Contributing and BUG reporting
https://github.com/jhcepas/ete/blob/master/CONTRIBUTING.rst
Getting Support
For any type of question on how to use ETE in bioinformatics, the BioStars community (http://biostars.org) provides an excellent help desk. ETE developers contribute there with answers, but will also get feedback from other users. It is recommended to tag your questions with the "etetoolkit" label.
For technical problems or more ETE specific questions, you can also use the official ETE mailing list at https://groups.google.com/d/forum/etetoolkit. To avoid spam, messages from new users are moderated. Expect some delay until your first message and account is validated.
As this mailing list is regularly checked for pending messages by the ETE developers, messages are more likely to be addressed than direct support questions sent to the developers' emails.
Bug reports and feature requests are usually discussed at https://github.com/etetoolkit/ete/issues
For any other inquire, preferably non-support questions, please contact huerta /at/ embl.de