Comments (5)
Check this issue #37 .Need to update the tutorial.
from lightdock.
Indeed, this is a problem with the tutorial, needs to be updated as @alchemistcai suggests.
from lightdock.
As a workaround, @zhl10154 please use -spr 10 during setup to fix the maximum number of swarms per restraints to 10 and having a similar number of swarms compared to the tutorial.
from lightdock.
As a workaround, @zhl10154 please use
-spr 10during setup to fix the maximum number of swarms per restraints to 10 and having a similar number of swarms compared to the tutorial.
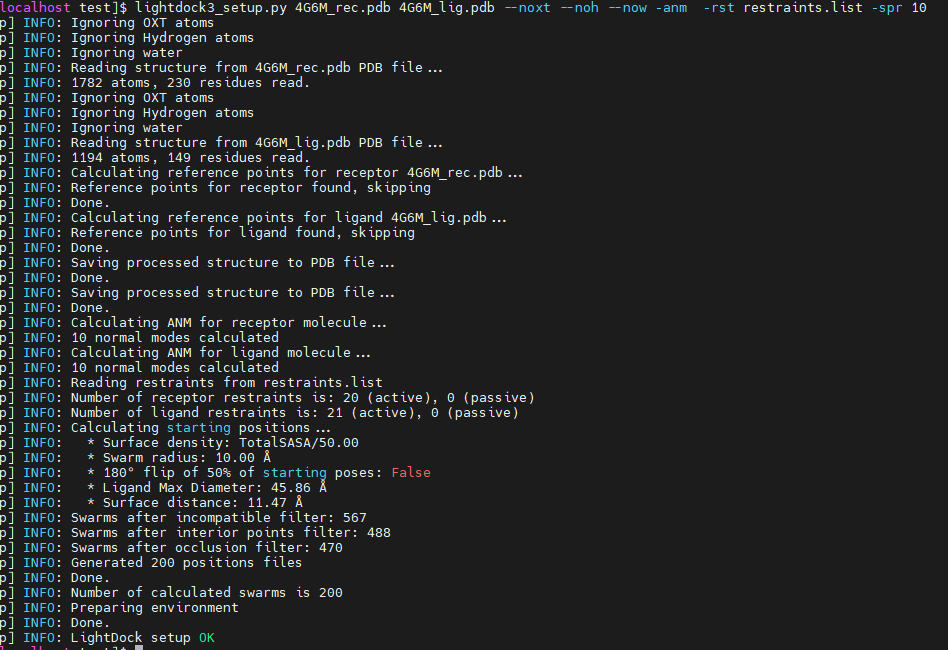
Thanks, man. I tried -spr 10, swarm number reduced to 200 (it might calculated by 10*20, which 20 is number of restraints), it still takes a lot of time to compute. Is there anyway to make it self-adaptive?
from lightdock.
It is self-adaptive to the number of restraints and region to sample. If you wish to decrease the number, just tweak -spr according to your expectations. In case you want to reproduce the current tutorial, please reinstall lightdock at version 0.9.2. If you want to speedup calculations, use the Rust implementation of LightDock.
from lightdock.
Related Issues (20)
- what the datas meaning of cluster.repr file? HOT 4
- Unexpected keyword argument 'seed' HOT 6
- dna.pdb file empty after running `reduce_to_amber.py` and subsequent error during simulation HOT 10
- lightdock pdb file visualises incorrectly in pymol HOT 5
- ERROR: [NormalModesCalculationError] Number of atoms is different HOT 6
- Can't find implicit membrane model for PBP2a protein on Memdock. HOT 6
- Issues when docking HOT 1
- test_dna and test_pydock failed. HOT 2
- TOBI scoring function fails to replicate results with CCharPPI server HOT 5
- Error encountered with DNA scoring function in lightdock3.py for DNA-DNA docking HOT 3
- Migrate tests from nose to pytest HOT 2
- Question about the treatment of protein-ligand docking. HOT 3
- Freezes after a while HOT 6
- Support for non standard amino acids HOT 2
- Ligands are not placed in designated swarm centers HOT 6
- Performance for protein-DNA docking HOT 1
- Use MPI for simulation HOT 3
- Much larger number of swarms in restraints tutorial than shown in docs HOT 4
- [lgd_cluster_bsas.py] Clustering has failed. HOT 9
Recommend Projects
-
 React
React
A declarative, efficient, and flexible JavaScript library for building user interfaces.
-
Vue.js
🖖 Vue.js is a progressive, incrementally-adoptable JavaScript framework for building UI on the web.
-
 Typescript
Typescript
TypeScript is a superset of JavaScript that compiles to clean JavaScript output.
-
TensorFlow
An Open Source Machine Learning Framework for Everyone
-
Django
The Web framework for perfectionists with deadlines.
-
Laravel
A PHP framework for web artisans
-
D3
Bring data to life with SVG, Canvas and HTML. 📊📈🎉
-
Recommend Topics
-
javascript
JavaScript (JS) is a lightweight interpreted programming language with first-class functions.
-
web
Some thing interesting about web. New door for the world.
-
server
A server is a program made to process requests and deliver data to clients.
-
Machine learning
Machine learning is a way of modeling and interpreting data that allows a piece of software to respond intelligently.
-
Visualization
Some thing interesting about visualization, use data art
-
Game
Some thing interesting about game, make everyone happy.
Recommend Org
-
Facebook
We are working to build community through open source technology. NB: members must have two-factor auth.
-
Microsoft
Open source projects and samples from Microsoft.
-
Google
Google ❤️ Open Source for everyone.
-
Alibaba
Alibaba Open Source for everyone
-
D3
Data-Driven Documents codes.
-
Tencent
China tencent open source team.

from lightdock.