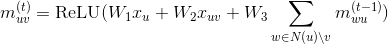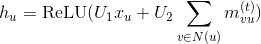Had a discussion with @ylfdq1118 today. Here is the conclusion we have reached:
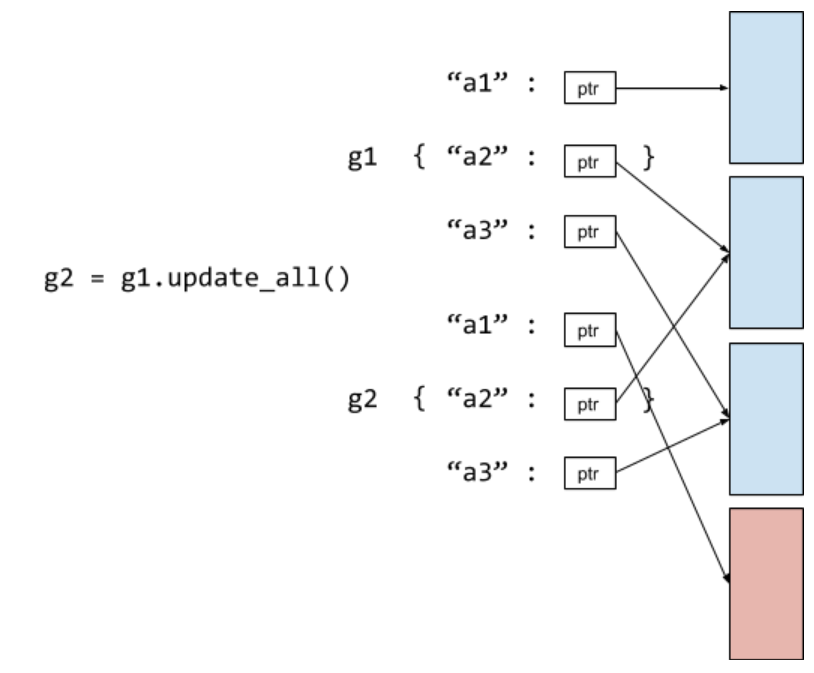
Storage
In our current codebase, we store the graph as adjacency lists (node to node). We propose to directly change the storage in C++ backend to incidence lists, more specifically, an incidence list for inbound edges and another for outbound ones. This implies that we are no longer handling simple graphs and multi-graphs separately like in NetworkX.
This will affect the implementation of the following backend methods:
pred and succ: Essentially, we will have to find the inbound/outbound edges of the given vertices first, then look up the source/target vertex of those edges (we can also have a predecessor/successor list in parallel to the inbound/outbound edges, but this is essentially duplicating the edge list and I'm not sure whether we would like it). These methods should support a boolean argument unique to dictate whether we expect a unique list of vertices (useful for graph traversal) or not (useful for deciding source/target nodes for message and apply_edge functions).vertex_subgraph: Similar to pred/succ, we need to do two-step queries.get_eid, in_edges, out_edges: Now it would return a list of edge IDs instead of one.get_eids, batch_in_edges, batch_out_edges: We haven't go through the semantics of this method yet. I would suggest returning a list of lists if possible, following the same practice as igraph.Graph.neighborhood(). Returning a padded array (along with the lengths) can also do the job if we wish to enable padding (which is a separate functionality we wish to consider I assume).(source, target, edge ID) tuples.
EDIT4: after all, the changes in C++ interface only involves changing the returning type of get_eid to IdArray and batch_get_eid to EdgeArray.
User Interface
Right now, the following user interfaces assumes that no multi-edge exist:
set_e_reprget_e_reprapply_edgessend (and consequently send_and_recv, pull, push, and update_all)
- Note that
pull, push and update_all only assumes that in the implementation due to usage of send; the interface itself is fine.
update_edge
Since the interfaces expects to take in and return tensors, we think that having multi-edges will complicate the semantics significantly for get_e_repr and set_e_repr in particular: the users often have to track of how many edges are between two given vertices (e.g. Given a multi-graph ((0, 1), (0, 1), (1, 2)), set_edge_repr([0, 1], [1, 2]) will actually require a tensor with 3 rows, quite unnatural).
EDIT2: Our suggestion is to
- For
get_e_repr and set_e_repr, simply raise an error if multi-edge exists between any of the given source/target pairs. We can have nicer semantics later.
- For other methods, simply send/apply on all edges in between.
- Have a counterpart working on edge IDs for
apply_edges, send, send_and_recv, and update_edge (e.g. sendon) as well (set_e_repr and get_e_repr already have their edge ID counterparts), to allow sending/applying on individual edges.
We concluded that switching to incidence matrices in the backend should not break compatibility on the current user interfaces.
Side note: Filtering vertices/edges
Since RGCN already involves edges with different "types", which are represented by integers, we can expect that later on we will have models requiring to perform graph operations on a "type" of node/edge at a time. To be even more general, the users can do stuff on a subset of nodes/edges satisfying a certain predicate. We think it will be nice to have a filtering function which returns a subset of given nodes/edges that satisfies a given predicate. For example:
# returns all edges which has the field t equal to 2
eids = g.filter_edges(ALL, lambda e: e['t'] == 2)
# returns the nodes within @nids that has a maximum component greater than 1
# equivalent to something like nids[g.get_n_repr()['h'][nids].max(1) > 1]
nids = g.filter_nodes(nids, lambda n: n['h'].max(1) > 1)
Of course, the signature does not have to be exactly like this.
gSPMV Optimization
One thing we have noticed is that, if we use incidence matrices in our storage, then builtin reduce functions can be always represented by a gSPMV between the inbound incidence matrix and the message tensors.
However, if the message function is also a builtin, we don't want to actually broadcast the messages onto edges since it will potentially take up much more space. So
- If both message and reduce functions are builtins, we still stick to gSPMV on adjacency matrix if possible.
- Otherwise, if the reduce function is a builtin, we can do gSPMV on inbound incidence matrix.
EDIT: gSPMV on adjacency matrix with multi-edges is more subtle. Since multi-edges are analogous to duplicate entries in COO sparse matrix, depending on the message and reduce function form the actual implementation can be different:
- For message function
copy_src/src_mul_edge and reduce function sum, on PyTorch we can still stick with existing SPMM multiplications by directly constructing the (weighted) adjacency matrix from edge lists; PyTorch will automatically coalesce duplicated entries by summing them up. I don't know if this is the case for MXNet and Tensorflow.
- For other cases, it is very likely that we need to write our own operators to coalesce the duplicate entries. For instance, if the reduce function is
max, the coalescing operation is no longer a sum but something like max pooling, so direct sparse tensor construction in PyTorch no longer works.
RGCN example
Under the proposed changes, we expect that the RGCN code can be greatly simplified, and our behavior could match the official implementations. Namely, the message function is simply a batched dot product between the source tensor and the weight matrices corresponding to the edge types, while the reduce function is just a sum (@ylfdq1118 correct me if I'm wrong):
def msg(src, edges, W):
# W is a 3D Tensor (n_types, in_features, out_features) inside a closure or a parameter in a PyTorch module
return src[:, None] @ W[edges['type']]
reducer = builtin.sumComments and questions are welcome.