chrischoy / fcgf Goto Github PK
View Code? Open in Web Editor NEWFully Convolutional Geometric Features: Fast and accurate 3D features for registration and correspondence.
License: MIT License
Fully Convolutional Geometric Features: Fast and accurate 3D features for registration and correspondence.
License: MIT License
Hello Chris,
by trying to run FCGF in a docker and face some error that i cant solve.
I am using the following script to build the container:
FROM nvidia/cuda:10.2-devel
RUN apt update \
&& apt install -y python3.7 python3-pip git \
&& python3.7 -m pip install --upgrade --force pip
# Required by ME
RUN apt install -y libopenblas-dev python3.7-dev
# ME installation requires this.
# Basicly just to use pip instead of pip3 ?
RUN ln -s /usr/bin/python3.7 /usr/bin/python && ln -s /usr/bin/pip3 /usr/bin/pip
# Keeps Python from generating .pyc files in the container
ENV PYTHONDONTWRITEBYTECODE=1
# Turns off buffering for easier container logging
ENV PYTHONUNBUFFERED=1
RUN apt-get install -y\
build-essential \
apt-utils \
ca-certificates \
wget \
git \
vim \
libssl-dev \
curl \
unzip \
unrar
# Torch versions: https://pytorch.org/get-started/previous-versions/
RUN python3.7 -m pip install torch==1.5.1 torchvision==0.6.1
RUN export CXX=g++-7
RUN pip install git+https://github.com/NVIDIA/[email protected]
RUN apt-get install -y libsm6 libxrender1 libfontconfig1 libpython3.7-dev libopenblas-dev
RUN apt install libgl1-mesa-glx -y
WORKDIR /app
# Avoid adding files by copy or add -> changes need new docker build
# use volumes
# only for intsallation purpose:
COPY ./FCGF/requirements.txt /tmp/FCGF/requirements.txt
RUN python3.7 -m pip install -r /tmp/FCGF/requirements.txt
CMD [ "/bin/bash", "" ]
and run it with the following script:
SCRIPT=$(readlink -f "$0")
SCRIPTPATH=$(dirname "$SCRIPT")
docker build -t mke_docker:0.5 .
docker rm -f mke0
docker run --privileged -it \
-v /media/wboschmann/Work-Drive2/SRS-WORK/Data/:/app/Data \
-v $SCRIPTPATH/FCGF:/app/FCGF \
--name mke0 \
--gpus all \
--entrypoint /bin/bash \
mke_docker:0.5
I tested by running the train skript for threedmatch and run into the following error
root@743a96027975:/app/FCGF# python train.py --threed_match_dir /app/Data/threedmatch/
12/17 18:07:00 ===> Configurations
12/17 18:07:00 out_dir: outputs
12/17 18:07:00 trainer: HardestContrastiveLossTrainer
12/17 18:07:00 save_freq_epoch: 1
12/17 18:07:00 batch_size: 4
12/17 18:07:00 val_batch_size: 1
12/17 18:07:00 use_hard_negative: True
12/17 18:07:00 hard_negative_sample_ratio: 0.05
12/17 18:07:00 hard_negative_max_num: 3000
12/17 18:07:00 num_pos_per_batch: 1024
12/17 18:07:00 num_hn_samples_per_batch: 256
12/17 18:07:00 neg_thresh: 1.4
12/17 18:07:00 pos_thresh: 0.1
12/17 18:07:00 neg_weight: 1
12/17 18:07:00 use_random_scale: False
12/17 18:07:00 min_scale: 0.8
12/17 18:07:00 max_scale: 1.2
12/17 18:07:00 use_random_rotation: True
12/17 18:07:00 rotation_range: 360
12/17 18:07:00 train_phase: train
12/17 18:07:00 val_phase: val
12/17 18:07:00 test_phase: test
12/17 18:07:00 stat_freq: 40
12/17 18:07:00 test_valid: True
12/17 18:07:00 val_max_iter: 400
12/17 18:07:00 val_epoch_freq: 1
12/17 18:07:00 positive_pair_search_voxel_size_multiplier: 1.5
12/17 18:07:00 hit_ratio_thresh: 0.1
12/17 18:07:00 triplet_num_pos: 256
12/17 18:07:00 triplet_num_hn: 512
12/17 18:07:00 triplet_num_rand: 1024
12/17 18:07:00 model: ResUNetBN2C
12/17 18:07:00 model_n_out: 32
12/17 18:07:00 conv1_kernel_size: 5
12/17 18:07:00 normalize_feature: True
12/17 18:07:00 dist_type: L2
12/17 18:07:00 best_val_metric: feat_match_ratio
12/17 18:07:00 optimizer: SGD
12/17 18:07:00 max_epoch: 100
12/17 18:07:00 lr: 0.1
12/17 18:07:00 momentum: 0.8
12/17 18:07:00 sgd_momentum: 0.9
12/17 18:07:00 sgd_dampening: 0.1
12/17 18:07:00 adam_beta1: 0.9
12/17 18:07:00 adam_beta2: 0.999
12/17 18:07:00 weight_decay: 0.0001
12/17 18:07:00 iter_size: 1
12/17 18:07:00 bn_momentum: 0.05
12/17 18:07:00 exp_gamma: 0.99
12/17 18:07:00 scheduler: ExpLR
12/17 18:07:00 icp_cache_path: /home/chrischoy/datasets/FCGF/kitti/icp/
12/17 18:07:00 use_gpu: True
12/17 18:07:00 weights: None
12/17 18:07:00 weights_dir: None
12/17 18:07:00 resume: None
12/17 18:07:00 resume_dir: None
12/17 18:07:00 train_num_thread: 2
12/17 18:07:00 val_num_thread: 1
12/17 18:07:00 test_num_thread: 2
12/17 18:07:00 fast_validation: False
12/17 18:07:00 nn_max_n: 500
12/17 18:07:00 dataset: ThreeDMatchPairDataset
12/17 18:07:00 voxel_size: 0.025
12/17 18:07:00 threed_match_dir: /app/Data/threedmatch/
12/17 18:07:00 kitti_root: /home/chrischoy/datasets/FCGF/kitti/
12/17 18:07:00 kitti_max_time_diff: 3
12/17 18:07:00 kitti_date: 2011_09_26
12/17 18:07:00 Loading the subset train from /app/Data/threedmatch/
12/17 18:07:00 Loading the subset val from /app/Data/threedmatch/
12/17 18:07:00 ResUNetBN2C(
(conv1): MinkowskiConvolution(in=1, out=32, region_type=RegionType.HYPER_CUBE, kernel_size=[5, 5, 5], stride=[1, 1, 1], dilation=[1, 1, 1])
(norm1): MinkowskiBatchNorm(32, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
(block1): BasicBlockBN(
(conv1): MinkowskiConvolution(in=32, out=32, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[1, 1, 1], dilation=[1, 1, 1])
(norm1): MinkowskiBatchNorm(32, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
(conv2): MinkowskiConvolution(in=32, out=32, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[1, 1, 1], dilation=[1, 1, 1])
(norm2): MinkowskiBatchNorm(32, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
)
(conv2): MinkowskiConvolution(in=32, out=64, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[2, 2, 2], dilation=[1, 1, 1])
(norm2): MinkowskiBatchNorm(64, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
(block2): BasicBlockBN(
(conv1): MinkowskiConvolution(in=64, out=64, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[1, 1, 1], dilation=[1, 1, 1])
(norm1): MinkowskiBatchNorm(64, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
(conv2): MinkowskiConvolution(in=64, out=64, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[1, 1, 1], dilation=[1, 1, 1])
(norm2): MinkowskiBatchNorm(64, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
)
(conv3): MinkowskiConvolution(in=64, out=128, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[2, 2, 2], dilation=[1, 1, 1])
(norm3): MinkowskiBatchNorm(128, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
(block3): BasicBlockBN(
(conv1): MinkowskiConvolution(in=128, out=128, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[1, 1, 1], dilation=[1, 1, 1])
(norm1): MinkowskiBatchNorm(128, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
(conv2): MinkowskiConvolution(in=128, out=128, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[1, 1, 1], dilation=[1, 1, 1])
(norm2): MinkowskiBatchNorm(128, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
)
(conv4): MinkowskiConvolution(in=128, out=256, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[2, 2, 2], dilation=[1, 1, 1])
(norm4): MinkowskiBatchNorm(256, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
(block4): BasicBlockBN(
(conv1): MinkowskiConvolution(in=256, out=256, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[1, 1, 1], dilation=[1, 1, 1])
(norm1): MinkowskiBatchNorm(256, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
(conv2): MinkowskiConvolution(in=256, out=256, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[1, 1, 1], dilation=[1, 1, 1])
(norm2): MinkowskiBatchNorm(256, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
)
(conv4_tr): MinkowskiConvolutionTranspose(in=256, out=128, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[2, 2, 2], dilation=[1, 1, 1])
(norm4_tr): MinkowskiBatchNorm(128, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
(block4_tr): BasicBlockBN(
(conv1): MinkowskiConvolution(in=128, out=128, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[1, 1, 1], dilation=[1, 1, 1])
(norm1): MinkowskiBatchNorm(128, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
(conv2): MinkowskiConvolution(in=128, out=128, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[1, 1, 1], dilation=[1, 1, 1])
(norm2): MinkowskiBatchNorm(128, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
)
(conv3_tr): MinkowskiConvolutionTranspose(in=256, out=64, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[2, 2, 2], dilation=[1, 1, 1])
(norm3_tr): MinkowskiBatchNorm(64, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
(block3_tr): BasicBlockBN(
(conv1): MinkowskiConvolution(in=64, out=64, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[1, 1, 1], dilation=[1, 1, 1])
(norm1): MinkowskiBatchNorm(64, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
(conv2): MinkowskiConvolution(in=64, out=64, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[1, 1, 1], dilation=[1, 1, 1])
(norm2): MinkowskiBatchNorm(64, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
)
(conv2_tr): MinkowskiConvolutionTranspose(in=128, out=64, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[2, 2, 2], dilation=[1, 1, 1])
(norm2_tr): MinkowskiBatchNorm(64, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
(block2_tr): BasicBlockBN(
(conv1): MinkowskiConvolution(in=64, out=64, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[1, 1, 1], dilation=[1, 1, 1])
(norm1): MinkowskiBatchNorm(64, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
(conv2): MinkowskiConvolution(in=64, out=64, region_type=RegionType.HYPER_CUBE, kernel_size=[3, 3, 3], stride=[1, 1, 1], dilation=[1, 1, 1])
(norm2): MinkowskiBatchNorm(64, eps=1e-05, momentum=0.05, affine=True, track_running_stats=True)
)
(conv1_tr): MinkowskiConvolution(in=96, out=64, region_type=RegionType.HYPER_CUBE, kernel_size=[1, 1, 1], stride=[1, 1, 1], dilation=[1, 1, 1])
(final): MinkowskiConvolution(in=64, out=32, region_type=RegionType.HYPER_CUBE, kernel_size=[1, 1, 1], stride=[1, 1, 1], dilation=[1, 1, 1])
)
12/17 18:07:02 Resetting the data loader seed to 0
Traceback (most recent call last):
File "train.py", line 84, in <module>
main(config)
File "train.py", line 63, in main
trainer.train()
File "/app/FCGF/lib/trainer.py", line 124, in train
val_dict = self._valid_epoch()
File "/app/FCGF/lib/trainer.py", line 321, in _valid_epoch
coordinates=input_dict['sinput0_C'].to(self.device))
File "/usr/local/lib/python3.7/dist-packages/MinkowskiEngine/MinkowskiTensor.py", line 327, in __init__
minkowski_algorithm=minkowski_algorithm,
File "/usr/local/lib/python3.7/dist-packages/MinkowskiEngine/MinkowskiCoordinateManager.py", line 142, in __init__
self._CoordinateManagerClass = getattr(_C, "CoordinateMapManager" + postfix)
AttributeError: module 'MinkowskiEngineBackend._C' has no attribute 'CoordinateMapManagerGPU_c10'
On the fist view everything seems fine, the systems seems to have cuda support and the required packeges seem to be available
root@743a96027975:/app/FCGF# pip list
Package Version
------------------- --------
argon2-cffi 20.1.0
asn1crypto 0.24.0
async-generator 1.10
attrs 20.3.0
backcall 0.2.0
bleach 3.2.1
cffi 1.14.4
cryptography 2.1.4
cycler 0.10.0
decorator 4.4.2
defusedxml 0.6.0
easydict 1.9
entrypoints 0.3
future 0.18.2
future-fstrings 1.2.0
idna 2.6
importlib-metadata 3.3.0
ipykernel 5.4.2
ipython 7.19.0
ipython-genutils 0.2.0
ipywidgets 7.5.1
jedi 0.17.2
Jinja2 2.11.2
joblib 1.0.0
jsonschema 3.2.0
jupyter-client 6.1.7
jupyter-core 4.7.0
jupyterlab-pygments 0.1.2
keyring 10.6.0
keyrings.alt 3.0
kiwisolver 1.3.1
MarkupSafe 1.1.1
matplotlib 3.3.3
MinkowskiEngine 0.5.0rc0
mistune 0.8.4
nbclient 0.5.1
nbconvert 6.0.7
nbformat 5.0.8
nest-asyncio 1.4.3
notebook 6.1.5
numpy 1.19.4
open3d 0.9.0.0
packaging 20.8
pandocfilters 1.4.3
parso 0.7.1
pexpect 4.8.0
pickleshare 0.7.5
Pillow 8.0.1
pip 20.3.2
prometheus-client 0.9.0
prompt-toolkit 3.0.8
protobuf 3.14.0
ptyprocess 0.6.0
pycparser 2.20
pycrypto 2.6.1
Pygments 2.7.3
pygobject 3.26.1
pyparsing 2.4.7
pyrsistent 0.17.3
python-dateutil 2.8.1
pyxdg 0.25
pyzmq 20.0.0
scikit-learn 0.23.2
scipy 1.5.4
SecretStorage 2.3.1
Send2Trash 1.5.0
setuptools 39.0.1
six 1.11.0
tensorboardX 2.1
terminado 0.9.1
testpath 0.4.4
threadpoolctl 2.1.0
torch 1.5.1
torchvision 0.6.1
tornado 6.1
traitlets 5.0.5
typing-extensions 3.7.4.3
wcwidth 0.2.5
webencodings 0.5.1
wheel 0.30.0
widgetsnbextension 3.5.1
zipp 3.4.0
Does anyone else face similar problems ?
I hope the problem is reproducable, if any futher information is requred just let me know.
Thanks in advance.
Hi Chris,
Very great work! I have been using your method for some time and it works very well.
I could have missed it and you have shared it somewhere. If that is not the case, is it possible that I can have the pre-trained Minkowski model on the 3dmatch dataset so that I can do some other comparisons that were not done in your original FCGF paper?
Thank you very much!
Xingtong
Thanks for publishing this great work! I made an initial experiment in which resunet model is trained on my custom dataset. Comparing to the 3DMatch and KITTI dataset, objects are pretty small in this dataset. For example, say the input is the point cloud of a mug, I would like to register the (partial) point cloud with corresponding CAD (complete) model points.
I was not able to get reasonable results. I noticed that the evaluation metrics used in your paper is too large for my dataset, e.g. \tau_1 = 0.1m (0.01m is suitable for my application). Am I doing something wrong or the FCGF model is actually not accurate enough to do object registration?
Dear Chris:
Thanks for your great job. I clone your FCGF code and run the train.py on my server. I run into a Segmentation Fault Error:

The checkpoint is set in /lib/data_loader.py which is :
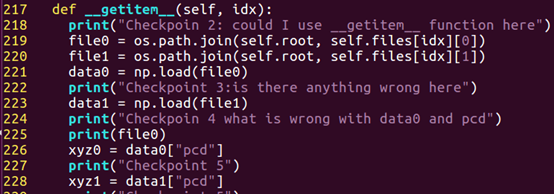
The server could not exec line “226” “xyz0=data0[“pcd”]”
But when I exec the data loading code in python:
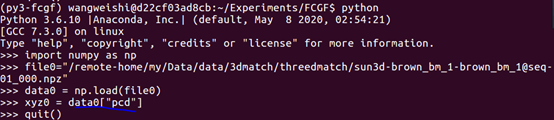
I have no problem like “Segmentation Error”.
And I am wondering what is wrong with this.
Best!
I cannot find the visualization_demo.py. Would you please upload it?
Hi, Thank you so much for your sharing. I found that the link to the model on Kitti dataset 20cm voxel size is missing. Could you please check it ?
Hi Chris,
I noticed that you only include voxels from overlapping volumes of point clouds. i.e only voxels that have at least one correspondence in the other cloud have a possibility to be included in the negative loss (and therefore the total loss).
neg_keys0 = _hash([pos_ind0.numpy(), D01ind], hash_seed)
...
mask0 = torch.from_numpy(
np.logical_not(np.isin(neg_keys0, pos_keys, assume_unique=False)))
...
neg_loss0 = F.relu(self.neg_thresh - D01min[mask0]).pow(2)
The problem arises when the overlapping volume of the given clouds is not that big: even when the loss is zero, on the validation voxels from non-overlapping areas can produce false positives that corrupt the registration (that is particularly strange to get a wrong result with the zero loss).
I tried to include negative matches from non-overlapping volume and it appears to fix the problem. What do you think about that? If you validate my idea I can propose apr for this issue.
Hello,
I have another question, and it is also about a memory problem.
When I proceed training.py, GPU memory problem occurred like this:
My GPU is GeForce GTX 1050 Ti, and I don't have a much better GPU.
How can I finish this training?
Is there any method to proceed with this task? (such as adjustment of a hyperparameter setting....)
Thank you in advance.
@chrischoy @sjnarmstrong ,Thanks for your sharing. I tried your code on 3DMatch dataset using the default configuration and found the training process is very slow. Specifically it took about one and a half hour for one epoch. (as you mentioned in the paper, you trained FCGF for 100 epochs, which means more than one week in my configuration). The GPU memory it took is only less than 5000 MB and GPU utility is less than 10% but CPU utility is high. I wonder is it normal situation and what's the most time-consuming part ?but I use V100 to train the model. And also find the speed of training on GTX1080Ti is faster than it on a V100.
In Issue#11, I could not find the solution, so can you provide another way to solve this problew
Thanks a lot.
Hello, thanks for sharing the well-organized code. I have downloaded the training data of 3dmatch from the link provided in scripts/download_datasets.sh. I found there only 7000+ scene pairs with 0.3 overlap threshold while there are tens of thousands frames in even one sequence of the original 3dmatch dataset. Have you conducted some sampling operations and are these all the training data?
Thanks in advance :)
In your latest commit, there is a bug in scripts/benchmark_3dmatch.py, line 175, saying:
However, there isn't any groundtruth log in testing dataset whose name ends with "_gt.log". After checking the testing dataset filename format, the code above will work if modified like this:
Maybe the testing dataset structure has been changed. Could you inspect it if possible?
Hello,
Thank you so much for sharing youse awesome work.
But I have a problem. When I tried to operate demo.py, I got the error like below:
with the black window like this (sometimes my computer is totally stuck with this situation):

For better understanding, my computer's spec is like this:

I think it has something to do with my memory usage, but I don't know at all why the error occurred.
Running demo.py script returns this error
Extension = ply
Traceback (most recent call last):
File "demo.py", line 73, in <module>
demo(config)
File "demo.py", line 41, in demo
skip_check=True)
File "/home/tpatten/Code/FCGF/util/misc.py", line 82, in extract_features
coords = coords[inds]
IndexError: shape mismatch: indexing arrays could not be broadcast together with shapes (20685,3) (20685,)```
Hi @chrischoy @sjnarmstrong
Thank you for sharing this wonderful work!
When I run the train.py script with 3dmatch dataset, something goes wrong, the error was report below:
09/26 12:59:16 Resetting the data loader seed to 0
Traceback (most recent call last):
File "train.py", line 84, in <module>
main(config)
File "train.py", line 63, in main
trainer.train()
File "/disk/tia/FCGF/lib/trainer.py", line 124, in train
val_dict = self._valid_epoch()
File "/disk/tia/FCGF/lib/trainer.py", line 314, in _valid_epoch
input_dict = data_loader_iter.next()
File "/home/ubuntu/.conda/envs/py3-fcgf/lib/python3.7/site-packages/torch/utils/data/dataloader.py", line 363, in __next__
data = self._next_data()
File "/home/ubuntu/.conda/envs/py3-fcgf/lib/python3.7/site-packages/torch/utils/data/dataloader.py", line 989, in _next_data
return self._process_data(data)
File "/home/ubuntu/.conda/envs/py3-fcgf/lib/python3.7/site-packages/torch/utils/data/dataloader.py", line 1014, in _process_data
data.reraise()
File "/home/ubuntu/.conda/envs/py3-fcgf/lib/python3.7/site-packages/torch/_utils.py", line 395, in reraise
raise self.exc_type(msg)
IndexError: Caught IndexError in DataLoader worker process 0.
Original Traceback (most recent call last):
File "/home/ubuntu/.conda/envs/py3-fcgf/lib/python3.7/site-packages/torch/utils/data/_utils/worker.py", line 185, in _worker_loop
data = fetcher.fetch(index)
File "/home/ubuntu/.conda/envs/py3-fcgf/lib/python3.7/site-packages/torch/utils/data/_utils/fetch.py", line 44, in fetch
data = [self.dataset[idx] for idx in possibly_batched_index]
File "/home/ubuntu/.conda/envs/py3-fcgf/lib/python3.7/site-packages/torch/utils/data/_utils/fetch.py", line 44, in <listcomp>
data = [self.dataset[idx] for idx in possibly_batched_index]
File "/disk/tia/FCGF/lib/data_loaders.py", line 252, in __getitem__
pcd0.colors = o3d.utility.Vector3dVector(color0[sel0])
IndexError: shape mismatch: indexing arrays could not be broadcast together with shapes (22361,3) (22361,)
Cloud you please help me to solve this problem?
Thank you very much!
Hi,
first of all thank you for sharing your code and for your inspiring work. I'm recently diving into it and I would like to ask you a question. I see that you perform a downsampling by dividing each point by the voxel size. Moreover, you claim in your paper that "As the input to the network requires unique coordinates C and corresponding features F, we first downsample the input point cloud". I'm missing the meaning of this and I cannot understand why you do such normalization of the data by dividing all points by the voxel size. If you can elaborate just a bit more this step it could be really helpful! Nevertheless, in the end we still have one 32-dimensional feature per point of our pointcloud or do we not? Thanks and sorry for my bad understanding!
Thanks again, and have a nice day
Marco
hi Chris,
In the contrastive loss implementation, for positive part, you use F.relu(squared_distance - positive_margin); for negative part, you use F.relu(negative_margin - squared_distance)**2, is this intentional?
Line 447 in 1a4e35c
best,
Shengyu
Hi, chrischoy. Thanks a lot for the nice work and code.
I have a question about processing unordered point sets.
In prior works like Pointnet, due to the unorderedness of point sets, Maxpooling is usually used to promise the stability of extracted descriptors, but I do not find any description about the unordered point sets in the paper. Could you offer me some help?
The script visualization_demo.py does not seem to be in the repository.
Can you check?
Thank you!
How can I create GT trans parameters of my own ply files?
I get a result with different color in CUDA11.1+cudatoolkit 11.0.2+pytorch1.7.1
the link to the dataset seems broken.
First of all, thanks for sharing the code. This is an amazing project. Now I meet some problem.
I do some test on real-world data using the default weight(which are downloaded from here). The result seems bad. The colors in the same places should be the same, but they are very different.

What I do to the colored mesh is a voxel clustering simplification, because I need to download it to see. And I don't think this influences the visualization result. Is the defualt weight not right for general scene? Should I retrain the model using kitti dataset?
Thanks again!
Hi Chris,
I saw in your code, sample_random_trans() use np.pi / 4 as the input argument in some dataset classes. However, according to the function definition, it actually should accept the unit of degree instead of radian as the argument. I am afraid this may affect your experiment results in your paper if this is actually the case.
Hi Chris,
Thank you for sharing the code!
I wonder which algorithm is using here for transformation estimation in the util/transform_estimation.py. Can you point out the reference?
Thanks!
Qiaojun
First off, thank you for open sourcing this code! It's readable and very helpful; I loved the paper and found the results to be very exciting!
I was able to kick off an evaluation of the KITTI results using the model "ResUNetBN2C, Normalization=False, KITTI, 20cm, 32-dim". Since the required config.json was not available, I reverse engineered one myself, which may be at least part of the issue I am having.
I am running the test_kitti.py script (modified to use an updated Open3d; I can provide a pull request soon!) using the aforementioned model and the config posted below.
The script works, and starts evaluating on 6.8k samples. The preliminary numbers look good, but evaluation is very slow. The feature computation time is ~400ms / sample, but the mean RANSAC time sits at about 40 seconds / sample. This seems very large considering it's saturating my 24-core Intel Xeon E5-2687W at ~99% for the entire duration.
Sample script output so far:
01/14 12:03:28 40 / 6857: Data time: 0.0050524711608886715
Feat time: 0.4292876958847046, Reg time: 36.63782391548157,
Loss: 0.041788674890995026, RTE: 0.03362400613997767, RRE: 0.001606260281446482, Success: 41.0 / 41 (100.0 %)
With this run time, evaluating all 6.8k test samples found by the script would take ~50 hours, which seems a lot.
I noticed that the RANSACConvergenceCriteria are the key knob to tune. Setting max_validations, the second argument, to something like 25 (instead of 10k) makes registration run in ~1s on my machine, but seems to deteriorate the RTE to ~11--12cm instead of 5--6cm. The success rate seems to remain unchanged.
test_kitti.py script to run with the latest Open3d (and maybe some extra comments I added while learning about it) be useful?Thank you,
Andrei
My system:
The config.json I "reverse engineered" to evaluate on KITTI:
{
"out_dir": "outputs/01_kitti_dummy_pretrained/",
"trainer": "HardestContrastiveLossTrainer",
"save_freq_epoch": 1,
"batch_size": 4,
"val_batch_size": 1,
"use_hard_negative": true,
"hard_negative_sample_ratio": 0.05,
"hard_negative_max_num": 3000,
"num_pos_per_batch": 1024,
"num_hn_samples_per_batch": 256,
"neg_thresh": 1.4,
"pos_thresh": 0.1,
"neg_weight": 1,
"use_random_scale": false,
"min_scale": 0.8,
"max_scale": 1.2,
"use_random_rotation": false,
"rotation_range": 360,
"train_phase": "train",
"val_phase": "val",
"test_phase": "test",
"stat_freq": 40,
"test_valid": true,
"val_max_iter": 400,
"val_epoch_freq": 1,
"positive_pair_search_voxel_size_multiplier": 1.5,
"hit_ratio_thresh": 0.1,
"triplet_num_pos": 256,
"triplet_num_hn": 512,
"triplet_num_rand": 1024,
"model": "ResUNetBN2C",
"model_n_out": 32,
"conv1_kernel_size": 7,
"normalize_feature": false,
"dist_type": "L2",
"best_val_metric": "feat_match_ratio",
"optimizer": "SGD",
"max_epoch": 100,
"lr": 0.1,
"momentum": 0.8,
"sgd_momentum": 0.9,
"sgd_dampening": 0.1,
"adam_beta1": 0.9,
"adam_beta2": 0.999,
"weight_decay": 0.0001,
"iter_size": 1,
"bn_momentum": 0.05,
"exp_gamma": 0.99,
"scheduler": "ExpLR",
"icp_cache_path": "/home/andreib/.cache/fcgf_icp_cache_path",
"use_gpu": true,
"weights": null,
"weights_dir": null,
"resume": null,
"resume_dir": null,
"train_num_thread": 2,
"val_num_thread": 1,
"test_num_thread": 2,
"fast_validation": false,
"nn_max_n": 500,
"dataset": "KITTIPairDataset",
"voxel_size": 0.20,
"threed_match_dir": "/home/chrischoy/datasets/FCGF/threedmatch",
"kitti_root": "<my kitti root>",
"kitti_max_time_diff": 3,
"kitti_date": "2011_09_26"
}
Dear Chris,
I took a look at your training data from this link: https://node1.chrischoy.org/data/datasets/threedmatch.tgz
This is a point cloud set with colors encoded. However, on the 3DMatch website. It only provides 2d RGB of these scenes. Could you please tell me what's preprocessing from 2d RGB to 3d point cloud? Thank you very much!
hi Chris, I open this issue regarding the generalisation ability of FCGF on ETH dataset here: XuyangBai/D3Feat#1. Can you help check this?
Dear Chris,
When I finished training and tried to evaluate the result following your instruction. It gives an error on mean of empty slice.
Full Error Message:
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
02/24 20:37:01 0.100000 0.050000
/disk_1/FCGF/scripts/benchmark_3dmatch.py:182: RuntimeWarning: Mean of empty slice.
logging.info("average : %.4f +- %.4f" % (scene_r.mean(), scene_r.std()))
/disk_1/anaconda3/envs/fcgf/lib/python3.7/site-packages/numpy/core/_methods.py:161: RuntimeWarning: invalid value encountered in double_scalars
ret = ret.dtype.type(ret / rcount)
/disk_1/anaconda3/envs/fcgf/lib/python3.7/site-packages/numpy/core/_methods.py:217: RuntimeWarning: Degrees of freedom <= 0 for slice
keepdims=keepdims)
/disk_1/anaconda3/envs/fcgf/lib/python3.7/site-packages/numpy/core/_methods.py:186: RuntimeWarning: invalid value encountered in true_divide
arrmean, rcount, out=arrmean, casting='unsafe', subok=False)
/disk_1/anaconda3/envs/fcgf/lib/python3.7/site-packages/numpy/core/_methods.py:209: RuntimeWarning: invalid value encountered in double_scalars
ret = ret.dtype.type(ret / rcount)
02/24 20:37:01 average : nan +- nan
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
Command input for running:
python -m scripts.benchmark_3dmatch --source /disk_1/FCGF/3DMatch/threedmatch/ --target ./feature_tmp/ --voxel_size 0.025 --model /disk_1/FCGF/outputs/checkpoint.pth --do_generate --do_exp_feature --with_cuda
Do you have any idea on this error? Dataset is downloaded through command:
./scripts/download_datasets.sh /path/to/dataset/download/dir
And training is perfectly fine.
Hi Chris,
Thanks for your work !
I just have a little question.
In your article, you said that the hardest triplet loss is prone to collapse... To mitigate the problem, you mix hardest triplet with randomly sampled triplet.
Do you know why is it prone to collapse ?
Dear Chris,
In Leonhard cluster, during code run, an error occurs which I didn't come across when running code locally. Specifically in this line:https://github.com/StanfordVL/MinkowskiEngine/blob/a0a8ce3ac6aab1e3107a057ff9ec3394087ade6c/MinkowskiEngine/MinkowskiConvolution.py#L86.
I have also attached the full log for this error.
lsf.o5456313.txt
Dear Chris,
Due to my limited GPU memory, I was wondering if MinkowskiEngine support float16 computation? I am getting an error when making model as model.to(dtype=tensor.float16), then I am getting this error.
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
f"Type mismatch input: {input_features.type()} != kernel: {kernel.type()}"
AssertionError: Type mismatch input: torch.cuda.FloatTensor != kernel: torch.cuda.HalfTensor
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
Is that maybe because your pretrained model is in float32?
Thank you in advance.
Thanks for your work!
I have run the demo code and visualized the features. However, I don't know how to test with kitti odometry data. Could you give some instructions for running /scrips/test_kitti ?
Hi @chrischoy @sjnarmstrong
Thanks for sharing your wonderful projects!
I comment the line252-255
Line 252 in 1a4e35c
feats0:(array([], shape=(0, 1), dtype=float64),)
Traceback (most recent call last):
File "train.py", line 84, in <module>
main(config)
File "train.py", line 63, in main
trainer.train()
File "/disk/tia/FCGF/lib/trainer.py", line 124, in train
val_dict = self._valid_epoch()
File "/disk/tia/FCGF/lib/trainer.py", line 314, in _valid_epoch
input_dict = data_loader_iter.next()
File "/home/ubuntu/.conda/envs/py3-fcgf/lib/python3.7/site-packages/torch/utils/data/dataloader.py", line 363, in __next__
data = self._next_data()
File "/home/ubuntu/.conda/envs/py3-fcgf/lib/python3.7/site-packages/torch/utils/data/dataloader.py", line 989, in _next_data
return self._process_data(data)
File "/home/ubuntu/.conda/envs/py3-fcgf/lib/python3.7/site-packages/torch/utils/data/dataloader.py", line 1014, in _process_data
data.reraise()
File "/home/ubuntu/.conda/envs/py3-fcgf/lib/python3.7/site-packages/torch/_utils.py", line 395, in reraise
raise self.exc_type(msg)
AssertionError: Caught AssertionError in DataLoader worker process 0.
Original Traceback (most recent call last):
File "/home/ubuntu/.conda/envs/py3-fcgf/lib/python3.7/site-packages/torch/utils/data/_utils/worker.py", line 185, in _worker_loop
data = fetcher.fetch(index)
File "/home/ubuntu/.conda/envs/py3-fcgf/lib/python3.7/site-packages/torch/utils/data/_utils/fetch.py", line 47, in fetch
return self.collate_fn(data)
File "/disk/tia/FCGF/lib/data_loaders.py", line 71, in collate_pair_fn
coords_batch0, feats_batch0 = ME.utils.sparse_collate(coords0, feats0)
File "/home/ubuntu/.conda/envs/py3-fcgf/lib/python3.7/site-packages/MinkowskiEngine/utils/collation.py", line 124, in sparse_collate
assert N == Nf, f"Coordinate length {N} != Feature length {Nf}"
AssertionError: Coordinate length 135925 != Feature length 0
It seems that that data was failed to extract in
Line 27 in 1a4e35c
By the way, I test the same 3dmatch dataset on Deep Global Registration, there is no error during training, it proves that the dataset is downloaded correctly.
Looking forward to your reply!
In the paper, the percentage of overlap is at least 30%. But I was not able to get the training to converge on my dataset which has 50% overlaps. The input point clouds are from one CAD model, one point cloud keeps all the sampled surface points from CAD, the other one is obtained by first cuts the CAD model into half, and then sampled surface points from half of the model.
Hi, im trying to run demo.py after folllowing the installation instaructions. i get:
File "demo.py", line 73, in
demo(config)
File "demo.py", line 29, in demo
model = ResUNetBN2C(1, 16, normalize_feature=True, conv1_kernel_size=3, D=3)
File "/home/ubuntu/FCGF/model/resunet.py", line 38, in init
dimension=D)
TypeError: init() got an unexpected keyword argument 'bias'
As shown here:
The newer versions of ME make use of *sparce_tensors as opposed to a single tuple value.
The current models make use of the tuple definition
Hi @chrischoy Thanks for your sharing. I tried your code on 3DMatch dataset using the default configuration and found the training process is very slow. Specifically it took about one and a half hour for one epoch. (as you mentioned in the paper, you trained FCGF for 100 epochs, which means more than one week in my configuration). The GPU memory it took is only less than 5000 MB and GPU utility is less than 10% but CPU utility is high. I wonder is it normal situation and what's the most time-consuming part ? I use RTX 2080Ti to train the model.
Thanks a lot.
Hi @chrischoy , I think this line should be
coords = np.hstack([np.zeros((len(coords), 1)), coords])
# coords = np.hstack([coords, np.zeros((len(coords), 1))])as you have updated the MinkowskiEngine to version 0.4
I get a result with different color in CUDA11.1+cudatoolkit 11.0.2+pytorch1.7.1
Hello.
It was really helpful to understand about Minkowski engine and feature extraction of 3D bodies.
I am really interested to know more about feature extraction.
Will the feature extraction speed get affected if we use keras also into the code along with Minkowski engine.
Hi thanks for the amazing work!
During the evaluation of pairwise registration, your code seems to compute the nearest neighbour in the second point cloud for each point in the first point cloud. Does that mean that the number of correspondences your model produced is equal to the number of points in the first point cloud?
If so, how do you handle the points with no correspondences when you are computing the 'Feature-Match recall' metric?
Hello, i have this problem where i run out of memory when running python train.py --threed_match_dir ~/dataset/threedmatch/ --batch_size 1.
At first i ran out of memory before even starting the first epochs, so i changed the batch_size to 1 (batch_size 2 was still too much). After going through some thousands epochs i started getting "out of memory" errors like:
INFO - 2021-02-22 12:51:28,348 - trainer - Train Epoch: 1 [1440/7317], Current Loss: 1.157e+00 Pos: 0.365 Neg: 0.792 Data time: 0.0536, Train time: 0.5614, Iter time: 0.6150
Traceback (most recent call last):
File "train.py", line 84, in <module>
main(config)
File "train.py", line 63, in main
trainer.train()
File "/home/f/repos/FCGF/lib/trainer.py", line 132, in train
self._train_epoch(epoch)
File "/home/f/repos/FCGF/lib/trainer.py", line 492, in _train_epoch
self.config.batch_size)
File "/home/f/repos/FCGF/lib/trainer.py", line 427, in contrastive_hardest_negative_loss
D01 = pdist(posF0, subF1, dist_type='L2')
File "/home/f/repos/FCGF/lib/metrics.py", line 24, in pdist
D2 = torch.sum((A.unsqueeze(1) - B.unsqueeze(0)).pow(2), 2)
RuntimeError: CUDA out of memory. Tried to allocate 32.00 MiB (GPU 0; 3.82 GiB total capacity; 744.27 MiB already allocated; 43.38 MiB free; 814.00 MiB reserved in total by PyTorch)
Currently i'm my system takes up 500MiB VRAM from my GTX 1650 (4GB) and the rest is used by pytorch. I'm running pytorch 1.7 in a python 3.7 conda enviroment and i tried compiling minkowskiEngine with cuda 11.2 and currently i'm running cuda 10.2 but both gave the same error.
On a side note: Isn't it bad to run a batch size of only 1, wouldn't that cause poor convergence?
Hi Chris,
it is not really an issue but when performing the validation you can wrap the code in
with torch.no_grad():
such that the gradients are not computed.
It helps reduce the memory consumption and makes the validation faster.
Cheers
Zan
Hi chrischoy, @chrischoy
Nice work and thank you for sharing the code.
I have a question about the implementation of ResUNet2. In my opinion, the forward() function of ResUNet2 should look like:
def forward(self, x):
out_s1 = self.conv1(x)
out_s1 = self.norm1(out_s1)
out_s1 = MEF.relu(out_s1)
out= self.block1(out_s1)
...As shown in the ResUNet architecture in the paper:

However, your implement of ResUNet2 feed output of residual block to 3D ConvTr, and activate feature map twice using ReLU(one in self.block, one in MEF.relu). Is there a problem here, or am I missing something?
Lines 142 to 147 in 458549e
A declarative, efficient, and flexible JavaScript library for building user interfaces.
🖖 Vue.js is a progressive, incrementally-adoptable JavaScript framework for building UI on the web.
TypeScript is a superset of JavaScript that compiles to clean JavaScript output.
An Open Source Machine Learning Framework for Everyone
The Web framework for perfectionists with deadlines.
A PHP framework for web artisans
Bring data to life with SVG, Canvas and HTML. 📊📈🎉
JavaScript (JS) is a lightweight interpreted programming language with first-class functions.
Some thing interesting about web. New door for the world.
A server is a program made to process requests and deliver data to clients.
Machine learning is a way of modeling and interpreting data that allows a piece of software to respond intelligently.
Some thing interesting about visualization, use data art
Some thing interesting about game, make everyone happy.
We are working to build community through open source technology. NB: members must have two-factor auth.
Open source projects and samples from Microsoft.
Google ❤️ Open Source for everyone.
Alibaba Open Source for everyone
Data-Driven Documents codes.
China tencent open source team.